By Kevin E. Noonan —
 For several years, Sigma Aldrich has been prosecuting several applications (including USSNs 15/188,911; 15/188,924; and 15/456,204) claiming CRISPR technology that (it alleged) would be deserving of an interference with University of California's U.S. Application Nos. 15/9547,718 and 15/981,809, and reserving the right to supplement its request to include other patents and patent applications owned by the University of California Berkeley et al. (collectively "CVC") as well as those of owned by The Broad Institute and colleagues (see "Sigma-Aldrich Wants Its Piece of CRISPR Pie" and "Sigma-Aldrich Tries Again"). Such an interference, if declared, threatened to upset the CVC/Broad apple cart regarding CRISPR-Cas-9 patent ownership, Sigma–Aldrich alleging an earlier priority date than the Broad's earliest date and 3-7 months after CVC's earliest date:
For several years, Sigma Aldrich has been prosecuting several applications (including USSNs 15/188,911; 15/188,924; and 15/456,204) claiming CRISPR technology that (it alleged) would be deserving of an interference with University of California's U.S. Application Nos. 15/9547,718 and 15/981,809, and reserving the right to supplement its request to include other patents and patent applications owned by the University of California Berkeley et al. (collectively "CVC") as well as those of owned by The Broad Institute and colleagues (see "Sigma-Aldrich Wants Its Piece of CRISPR Pie" and "Sigma-Aldrich Tries Again"). Such an interference, if declared, threatened to upset the CVC/Broad apple cart regarding CRISPR-Cas-9 patent ownership, Sigma–Aldrich alleging an earlier priority date than the Broad's earliest date and 3-7 months after CVC's earliest date:
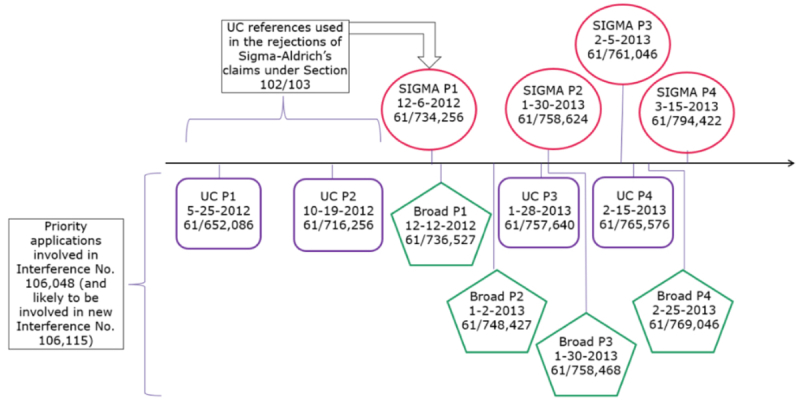
However, the U.S. Patent Examiner had taken the position that Sigma-Aldrich was not entitled to a Declaration of Interference until it had obtained allowed claims, and that CVC's earlier patent dates prevented such an allowance unless, inter alia, Sigma-Aldrich could swear behind the earlier patent application filing dates. With significant justification, Sigma-Aldrich characterized this situation as a Catch 22 in view of the seemingly contradictory conclusion arrived at by both Patent Trial and Appeal Board (see "PTAB Decides CRISPR Interference — No interference-in-fact" and "PTAB Decides CRISPR Interference in Favor of Broad Institute — Their Reasoning") and Federal Circuit (see "Regents of the University of California v. Broad Institute, Inc. (Fed. Cir. 2018)").
The logjam was recently broken for two of these Sigma-Aldrich applications, the Patent Office issuing a Notice of Allowance for 15/188,911 and 15/188,924. The allowed claims, including Examiner's amendments, are as follows:
U.S. Application No. 15/188911:
1. (currently amended) A method for integrating an exogenous sequence into a chromosomal sequence of a eukaryotic cell, the method comprising:
introducing into the eukaryotic cell:
(i) at least one RNA-guided endonuclease or nucleic acid encoding at least one RNA-guided endonuclease, wherein the at least one RNA-guided endonuclease is a clustered regularly interspersed short palindromic repeats (CRISPR)/CRISPR associated (Cas) (CRISPR-Cas) type II system protein, wherein the nucleic acid encoding the CRISPR-Cas type II system protein is codon optimized for expression in the eukaryotic cell,
wherein the CRISPR-Cas type II system protein is a Streptococcus pyogenes Cas9 protein including at least one nuclear localization signal consisting of comprising SEQ ID NO: 1 or SEQ ID NO:2 covalently attached to the C-terminal amino acid of the Cas9 protein sequence; and
(ii) at least one engineered guide RNA or DNA encoding at least one engineered guide RNA, each guide RNA comprising
(1) a first region at the 5' end that base pairs with a target site in the chromosomal sequence, and
(2) a second region that forms a secondary structure which interacts with the at least one RNA-guided endonuclease; and
(iii) at least one donor polynucleotide comprising the exogenous sequence;
whereby the at least one guide RNA guides the at least one RNA-guided endonuclease to the target site in the chromosomal sequence where the RNA-guided endonuclease introduces a double-stranded break, the target site in the chromosomal sequence is immediately followed by a proto spacer adjacent motif (PAM), and repair of the double-stranded break by a DNA repair process leads to integration of the exogenous sequence into the chromosomal sequence.
8. (original) The method of claim 1, wherein the exogenous sequence in the donor polynucleotide is flanked by sequences having substantial sequence identity to sequences on either side of the target site in the chromosomal sequence.
9. (previously presented) The method of claim 1, wherein the exogenous sequence in the donor polynucleotide further comprises a targeted cleavage site that is recognized by the at least one RNA-guided endonuclease.
10. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is mRNA
11. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is DNA
13. (previously presented) The method of claim 1, wherein the eukaryotic cell is a human cell, a nonhuman mammalian cell, or a plant cell.
14. (original) The method of claim 1, wherein the eukaryotic cell is in vitro.
15. (original) The method of claim 1, wherein the eukaryotic cell is in vivo.
16. (original) The method of claim 1, wherein the at least one guide RNA is at least partially chemically synthesized.
18. (previously presented) The method of claim [[19]] 1, wherein the Cas9 protein is from Streptococcus pyogenes strain MGAS15252 and comprises SEQ ID NO:9.
20. (currently amended) The method of claim [[19]] 1, wherein the at least one nuclear localization signal covalently attached to the C-terminal amino acid of the Cas9 protein sequence consists of comprises SEQ ID NO:1.
21. (currently amended) The method of claim 1, wherein the at least one RNA guided endonuclease comprising the at least one nuclear localization signal covalently attached to the C-terminal amino acid of the Cas9 protein sequence consists of comprises SEQ ID NO:2.
22. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is mRNA and the at least one guide RNA is comprised of two non-covalently bound RNA molecules.
23. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is mRNA and the at least one guide RNA is comprised of a single RNA molecule.
24. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is DNA, the at least one guide RNA is encoded by DNA, and the at least one guide RNA is comprised of a single RNA molecule.
25. (previously presented) The method of claim 1, wherein the at least one donor polynucleotide is double stranded DNA.
26. (previously presented) The method of claim 1, wherein the at least one donor polynucleotide is single stranded DNA.
The Notice of Allowance states that:
The [Examiner's] amendment of the claimed method to recite, "wherein the CRISPR-Cas type II system protein is a Streptococcus pyogenes Cas9 protein including at least one nuclear localization signal consisting of SEQ ID NO: 1 or SEQ ID NO:2 covalently attached to the C-terminal amino acid of the Cas9 protein sequence" obviates the rejections of record.
This claim has been narrowed to limit the nuclear localization sequence (NLS) to consist of the expressly recited sequence identified by SEQ ID Nos. 1 or 2, which were not disclosed in the prior art including CVC's earlier priority documents. And it is interesting to note that while the Office has asserted dozens of CRISPR patents and applications in earlier rejections, only one patent (and a scientific reference) are noted in the Examiner's Reasons for Allowance.
With regard to U.S. Application No. 15/188,924, the allowed claims are:
1. (currently amended) A method for modifying a chromosomal sequence in a eukaryotic cell, the method comprising:
introducing into the eukaryotic cell
(i) at least one RNA-guided endonuclease or nucleic acid encoding at least one RNA-guided endonuclease, wherein the at least one RNA-guided endonuclease is a clustered regularly interspersed short palindromic repeats (CRISPR)/CRISPR associated (Cas) (CRISPR-Cas) type II system protein,
wherein the nucleic acid encoding the CRISPR-Cas type II system protein is codon optimized for expression in the eukaryotic cell,
wherein the CRISPR-Cas type II system protein is a Streptococcus pyogenes Cas9 protein including at least one nuclear localization signal consisting of comprising SEQ ID NO: 1 or SEQ ID NO:2 covalently attached to the C-terminal amino acid of the Cas9 protein sequence; and
(ii) at least one engineered guide RNA or DNA encoding at least one engineered guide RNA, each guide RNA comprising
(1) a first region at the 5' end that base pairs with a target site in the chromosomal sequence, and
(2) a second region that forms a secondary structure which interacts with the at least one RNA-guided endonuclease; and, optionally,
(iii) at least one donor polynucleotide; [[and]]
whereby the at least one guide RNA guides the at least one RNA-guided endonuclease to the target site in the chromosomal sequence where the RNA-guided endonuclease introduces a double-stranded break, the target site in the chromosomal sequence is immediately followed by a proto spacer adjacent motif (PAM), and repair of the double-stranded break by a DNA repair process leads to modification of the chromosomal sequence.
8. (previously presented) The method of claim 1, wherein the optional donor polynucleotide comprises a donor sequence that has at least one nucleotide change relative to the chromosomal sequence near the target site in the chromosomal sequence.
9. (original) The method of claim 8, wherein the donor sequence is flanked by sequences having substantial sequence identity to sequences on either side of the target site in the chromosomal sequence.
10. (previously presented) The method of claim 8, wherein the donor sequence further comprises a targeted cleavage site that is recognized by the at least one RNA guided endonuclease.
11. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is mRNA
12. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is DNA
14. (previously presented) The method of claim 1, wherein the eukaryotic cell is a human cell, a nonhuman mammalian cell, or a plant cell.
15. (original) The method of claim 1, wherein the eukaryotic cell is in vitro.
16. (original) The method of claim 1, wherein the eukaryotic cell is in vivo.
17. (original) The method of claim 1, wherein the optional donor polynucleotide is not introduced into the eukaryotic cell, and repair of the double-stranded break by a non-homologous end-joining repair process results in inactivation of the chromosomal sequence.
18. (original) The method of claim 1, wherein the optional donor polynucleotide is introduced into the eukaryotic cell, and repair of the double-stranded break results in a change of at least one nucleotide in the chromosomal sequence.
19. (original) The method of claim 1, wherein the at least one guide RNA is at least partially chemically synthesized.
20. (original) The method of claim 1, wherein only (i) and (ii) are introduced into the eukaryotic cell.
21. (original) The method of claim 1, wherein (i), (ii), and (iii) are introduced into the eukaryotic cell.
23. (currently amended) The method of claim [[24]] 1, wherein the Cas9 protein is from Streptococcus pyogenes strain MGAS15252 and comprises SEQ 10 NO:9.
25. (currently amended) The method of claim [[24]] 1, wherein the at least one nuclear localization signal covalently attached to the C-terminal amino acid of the Cas9 protein sequence consists of comprises SEQ 10 NO:1.
26. (currently amended) The method of claim 1, wherein the at least one RNA guided endonuclease comprising the at least one nuclear localization signal covalently attached to the C-terminal amino acid of the Cas9 protein sequence consists of comprises SEQ ID NO:2.
27. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is mRNA and the at least one guide RNA is comprised of two non-covalently bound RNA molecules.
28. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is mRNA and the at least one guide RNA is comprised of a single RNA molecule.
29. (previously presented) The method of claim 1, wherein the nucleic acid encoding the at least one RNA-guided endonuclease is DNA, the at least one guide RNA is encoded by DNA, and the at least one guide RNA is comprised of a single RNA molecule.
30. (previously presented) The method of claim 1, wherein the optional donor polynucleotide is double stranded DNA
31. (previously presented) The method of claim 1, wherein the optional donor polynucleotide is single stranded DNA
As with the '911 application, the Notice of Allowance states:
The amendment of the claimed method to recite, "wherein the CRISPR-Cas type II system protein is a Streptococcus pyogenes Cas9 protein including at least one nuclear localization signal consisting of SEQ ID NO: 1 or SEQ ID NO:2 covalently attached to the C-terminal amino acid of the Cas9 protein sequence" obviates the rejections of record.
And the claims are equally limited to embodiments consisting of SEQ ID Nos. 1 or 2.
Also, the claims in U.S. Application No. 15/456,204 are not yet allowed but remain as subject matter for an interference under Rule 202:
1. A method for modifying a chromosomal sequence in a eukaryotic cell by integrating a donor sequence, the method comprising introducing into the eukaryotic cell: (i) a Clustered Regularly Interspersed Short Palindromic Repeats (CRISPR)/CRISPR-associated (Cas) (CRISPR-Cas) type II protein linked to at least one nuclear localization signal (NLS) or a nucleic acid encoding the CRISPR-Cas type II protein linked to at least one NLS, wherein the CRISPR-Cas type II protein is a Cas9 protein, and the nucleic acid encoding the CRISPR-Cas type II protein is codon optimized for expression in the eukaryotic cell; (ii) a guide RNA or DNA encoding the guide RNA, wherein the guide RNA comprises a first region that is complementary to a target site in the chromosomal sequence that is immediately followed by a protospacer adjacent motif, and a second region that interacts with the CRISPR-Cas type II protein; and (iii) a donor polynucleotide comprising the donor sequence;
wherein the guide RNA guides the CRISPR-Cas type II protein to the target site in the chromosomal sequence, the CRISPR-Cas type II protein introduces a double stranded break at the target site, and repair of the double-stranded break by a DNA repair process leads to integration or exchange of the donor sequence into the chromosomal sequence.
(These claims have not been subject to a limiting Examiner's amendment.) Sigma-Aldrich has withdrawn its Request under 37 C.F.R. § 41.202 for the PTAB to declare an interference with the patents and patent applications currently at issue between CVC and the Broad in Interference No. 106,115. Sigma-Aldrich has maintained its Petition to the Director that the PTAB should declare an interference involving the claims of the '204 application without requiring an allowance ("Sigma-Aldrich does not withdraw, and specifically maintains, the portions of its Renewed Petition and its Partial Withdrawal and Supplementation directed to Sigma-Aldrich's U.S. Patent App. No. 15/456,204 ("the Sigma '204 Application")."
That the allowed claims are exceedingly narrow is not the impediment it otherwise might be, because should the PTAB declare an interference (and Sigma-Aldrich be awarded priority of invention), the "prior art" that necessitated acceptance of these narrow claims would no longer preclude Sigma-Aldrich from obtaining claims commensurate with the scope of CRISPR-Cas9 disclosed in its applications. The allowed claims merely overcome the procedural requirement for having allowed claims; indeed, Sigma-Aldrich affirmatively asserts in its most recent Office Action responses prior to allowance that it intends to file continuations applications having claims that interfere with CVC and Broad's patents and applications. Interference motion practice, as has occurred between CVC and the Broad over the past nine months (and slated for culmination at Oral Hearing on May 18th) would be available to Sigma-Aldrich to craft the scope of the count and the claims corresponding thereto to its advantage.
There is no time frame for Sigma-Aldrich's petition in the '204 application to be considered much less granted, but allowance of these claims complicates the already complicated web of determining who really owns CRISPR technology.
 Unified Patents and IPWatchdog will be offering a webinar entitled "In Denial: The PTAB's 314(a) Discretion, General Plastics, and NHK Spring" on May 21, 2020 at 12:00 pm (EDT). Gene Quinn of IPWatchdog.com and Shawn Ambwani of Unified Patents will discuss recent changes to § 314, focusing on General Plastics, NHK Spring, the data, and practice tips around them, and also look to the factors in both petitioner's and patent owner's behavior that the Board is free to consider, analyze policy concerns, and recommend changes.
Unified Patents and IPWatchdog will be offering a webinar entitled "In Denial: The PTAB's 314(a) Discretion, General Plastics, and NHK Spring" on May 21, 2020 at 12:00 pm (EDT). Gene Quinn of IPWatchdog.com and Shawn Ambwani of Unified Patents will discuss recent changes to § 314, focusing on General Plastics, NHK Spring, the data, and practice tips around them, and also look to the factors in both petitioner's and patent owner's behavior that the Board is free to consider, analyze policy concerns, and recommend changes.








