By Kevin E. Noonan –
One of the characteristics of autumn is a harvest of vegetables in the squash family, including the ubiquitous pumpkin (equally famous for pumpkin pie and infamous for pumpkin spiced varieties of seemingly any food product). Appropriately, a recent report in the scientific journal Molecular Plant disclosed the genetic relationship between two squash family species, showing an interesting pattern of genomic co-existence that has persisted for about three million years.
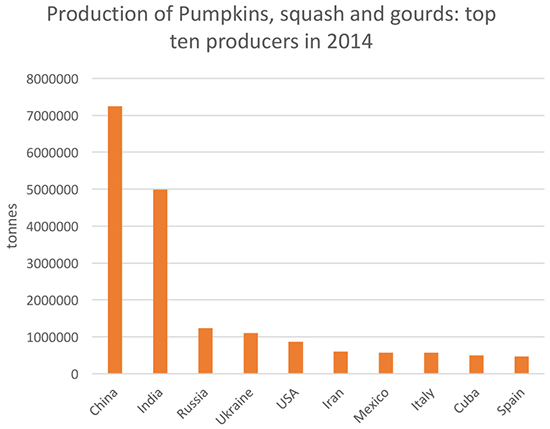
The report, entitled "Karyotype Stability and Unbiased Fractionation in the Paleo-Allotetraploid Cucurbita Genomes" was published by a group of researchers* from China and Cornell University in New York. Molecular Plant 10: 1293-306 (October 2017). The species, C. maxima Duch. and C. moschata Duch., are the most economically important members of the Cucurbitaceae family (Cucurbita species) having at least 10 wild and 5 domesticated species of squash. These squash species originated in South America (southern and northern regions, respectively) and were introduced into Europe following first contact. The species were secondarily domesticated in China/Japan and India/Myanmar, and today are cultivated globally (with annual production of 25.2 million tons for all species, predominantly in India and China; see chart below from "Pumpkin genomes sequenced revealing uncommon evolutionary history"). These species are also important as root stock for other produce (cucumbers, Cucumis sativus; watermelon, Citrullus lanatus; melon, Cucumis melo), and in particular one called Shintosa, a hybrid between C. maxima cv. Rimu and C. moschata cv. Rifu; C. maxima has better fruit flavor and texture but C. moschata has better disease resistance and resistance to abiotic stress.
Generically, Cucurbita species have a chromosome number of 40, which is higher than some other species, with evidence for whole-genome duplication being involved in evolution of this genus, but there was until now little whole genome sequence data available. Accordingly these authors describe whole genomic sequences for both species.
The results disclosed indicate an ancient allotetraploidization event that resulted in whole genome duplication, which the authors ascribe to "hybridization between two highly diverged diploid progenitors." The two genomes appear to have maintained the chromosomal structure of their progenitor species, having each experienced equivalent amounts of gene loss and neither of which showing dominance in gene expression levels (several homoeologous genes show divergent expression patterns). These data were used to analyze allele-specific gene expression in the Shintosa hybrid (F1).
The researchers sequenced the parent genomes of C. maxima cv. Rimu and C. moschata cv. Rifu at a total of 109.3 and 80.1 Gbps, respectively, giving an estimate of 386.6 and 371.0 Mkb respectively. Using 2030 SNPs the authors produced a draft genome of 271.4 Mb for C. maxima and 269.9 Mb C. moschata and constructed genetic map comprising 20 linkage groups (LGs)(corresponding to n=20 chromosomes in the haploid genome). These linkage groups were anchored to species-specific scaffolds, with 93% and 87% of the sequence reads mapping to the C. maxima and C. moschata assemblies, respectively. An assessment of the extent of repetitive sequences in the assembled genomes revealed about 40% in both, with about 70% of these repeat sequences being long terminal repeat (LTR) retrotransposons (mostly of the copia and gypsy type), consistent with results in other members of the family including cucumber, melon and watermelon.
mRNA sequences also mapped to these assemblies for C. maxima (93.8-95.8%) and C. moschata (86.2-95%). These RNA sequences mapped to an estimated 32,076 (C. maxima) and 32,205 (C. moschata) protein-coding genes, a much higher number than in cucumber (23,248 genes), melon (27,230 genes) or watermelon (23,440 genes). Sequence comparisons identified the biological functions of about 84% of the genes from each species, in which about 10% of the genes were identified as transcription factors. These analyses also identified disease-resistance genes; while most of these genes in C. maxima had homologues in C. moschata there were also disease-resistance genes that were unique to C. moschata.
Further comparisons of syntenic blocks in both C. maxima and C. moschata "clearly indicated" that both species underwent whole-genome duplication (WGD) also observed in some other Cucurbita species including pumpkin (C. pepo) but not others. The syntenic arrangement of the genes showed they corresponded to two subgenomes that could be distinguished according to genetic distance to orthologs for other related species (again, cucumber, melon and watermelon comprising the comparator set of related species). The authors drew the following conclusions:
Based on these findings, we separated the entire chromosomes into two groups to represent the two paleo-subgenomes. Eight and 11 chromosomes were assigned to subgenomes A and B, respectively. Chromosome 4 was divided into three segments with two assigned to subgenome A and one to subgenome B. The reconstructed subgenomes A and B of C. maxima contained 14,816 and 16,068 protein-coding genes, respectively (15,136 and 16,473 genes in C. moschata, respectively). Phylogenetic analysis clearly indicated an allotetraploid origin of Cucurbita from two progenitors that diverged from one another approximately 30.75 million years ago (Mya). One ancestor gave rise to watermelon, cucumber, and melon in the Benincaseae tribe, as well as to the B-genome progenitor of Cucurbita, which has not been sampled and may be extinct. No extant diploid species that could represent the A-genome progenitor have been sampled, so it is possible that this genome donor is also extinct. The oldest possible date for the polyploidy event is the date of divergence of Benincaseae from the B-genome of Cucurbita (approximately 26.28 Mya. There was extensive shared synteny between the two Cucurbita species with only a few inversions, consistent with a shared polyploidy event.
The report also assessed gene families amongst C. maxima, C. moschata, cucumber, melon, watermelon, grape, and papaya, and found 52 gene families that had expanded "significantly" during the natural history of these species. These included genes for pectin lyases, pathogenesis-related 1 proteins, carbohydrate esterases, pectinesterase inhibitors, subtilisin-like serine protease inhibitors, and mitogen-activated protein kinases (all of which are thought to act in modifying plant cell walls during growth and when fruit softens as well as in plant defense). The authors also reported expansions in gene families involved in meristem activity control and organ morphogenesis such as the gibberellin 3β-hydroxylase family. Although most of these expansions were found in both subgenomes, certain of them (such as pectin lyase and terpenene synthase) were from one subgenome while others (pectinesterase and carbohydrate esterase genes) were from the other subgenome.
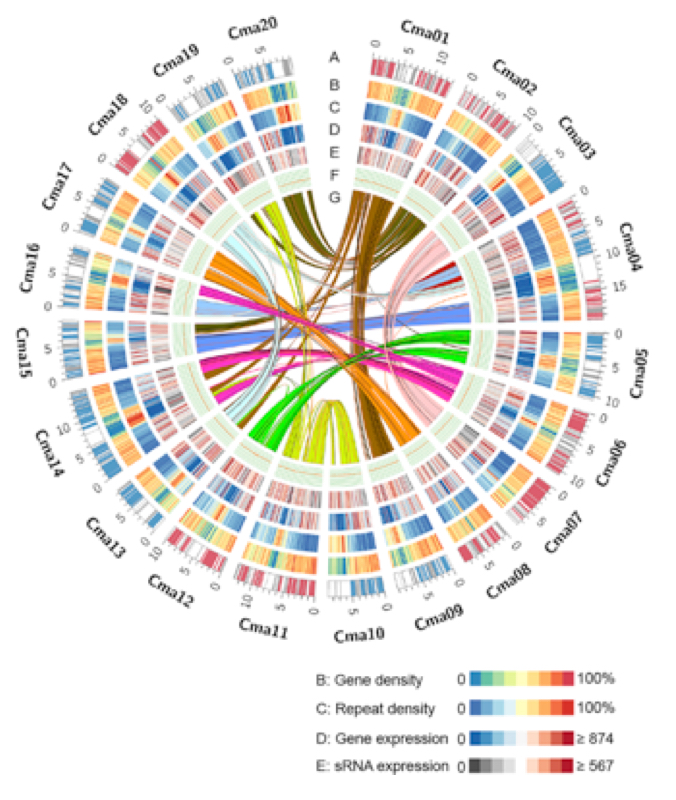
Figure 1 of Sun et al., 2017, Molecular Plant 10: 1293-306.
Another aspect of similarities and differences between C. maxima and C. moschata was divergence in gene expression between the two species, wherein between 22-32% of the genes were differentially expressed (distributed as 7303 in fruit, 6464 in leaf, 7749 in stem, and 9371 in root). Comparison of the mode of gene regulation (cis or trans) indicated that trans regulation was the predominant form of expression divergence in these species. This pattern was consistent, according to the authors, with a pattern of gene expression regulation evinced by less divergent species, in accord with their relatively recent evolutionary split. The report concludes with relative gene expression levels found in the F1 Shintosa hybrid, wherein changes in gene expression in root indicated a "trade off" between growth and defense responses, and the hybrid showed greater expression of genes involved in carotenoid biosynthesis.
The report contains a closing discussion of the unusual result that the two subgenomes appear for have engaged in extremely limited interchromosomal rearrangements since the hybridization of the two diploid progenitors that produced these species at least 3 million years ago. This pattern of chromosomal behavior has not been observed in other plant polyploidy species but has been observed in an allotetraploid frog with a long history as an experimental animal (Xenopus laevis); the hybridization event that created that species arose 17-18 million years ago.
* Honghe Sun (National Engineering Research Center for Vegetables, Beijing Academy of Agriculture and Forestry Sciences, Key Laboratory of Biology and Genetic Improvement of Horticultural Crops (North China), Beijing Key Laboratory of Vegetable Germplasm Improvement, Beijing, China and Boyce Thompson Institute, Cornell University, Ithaca, NY USA), Shan Wu (Boyce Thompson Institute), Guoyu Zhang (Beijing Key Laboratory of Vegetable Germplasm Improvement), Chen Jioa (Boyce Thompson Institute), Shaogui Guo (Beijing Key Laboratory of Vegetable Germplasm Improvement), Yi Ren (Beijing Key Laboratory of Vegetable Germplasm Improvement), Jie Zhang (Beijing Key Laboratory of Vegetable Germplasm Improvement), Haiying Zhang (Beijing Key Laboratory of Vegetable Germplasm Improvement), Guoyi Gong (Beijing Key Laboratory of Vegetable Germplasm Improvement), Zhangcai Jia (Beijing Key Laboratory of Vegetable Germplasm Improvement), Fan Zhang (Beijing Key Laboratory of Vegetable Germplasm Improvement), Jiaxing Tian (Beijing Key Laboratory of Vegetable Germplasm Improvement), William J. Lucas (Department of Plant Biology, College of Biological Sciences, University of California, Davis, CA, USA), Jeff J. Doyle (Section of Plant Breeding & Genetics, School of Integrated Plant Sciences, Cornell University), Haizhen Li (Beijing Key Laboratory of Vegetable Germplasm Improvement), Zhangjun Fe (Boyce Thompson Institute and USDA-ARS Robert W. Holley Center for Agriculture and Health, Ithaca, NY USA), and Yong Xu (Beijing Key Laboratory of Vegetable Germplasm Improvement).

Leave a comment