By Kevin E. Noonan —
 The rise and spread of the Zika virus, although limited for now to South America, has raised concerns globally, particularly with the prospect of the Summer Olympics in Rio de Janeiro later this year. And with those concerns comes renewed interest in the insect vector associated with zika virus transmission, the Aedes aegypti mosquito. This mosquito is also the transmission vector for other arboviruses, including those responsible for yellow fever, dengue fever, and Chikungunya, and has been extensively studied for over fifty years. These studies have been facilitated by the existence of a form of the mosquito that is readily adapted to the laboratory environment (perhaps related to the feral and domestic forms of the mosquito that exist in Africa).
The rise and spread of the Zika virus, although limited for now to South America, has raised concerns globally, particularly with the prospect of the Summer Olympics in Rio de Janeiro later this year. And with those concerns comes renewed interest in the insect vector associated with zika virus transmission, the Aedes aegypti mosquito. This mosquito is also the transmission vector for other arboviruses, including those responsible for yellow fever, dengue fever, and Chikungunya, and has been extensively studied for over fifty years. These studies have been facilitated by the existence of a form of the mosquito that is readily adapted to the laboratory environment (perhaps related to the feral and domestic forms of the mosquito that exist in Africa).
So it seems appropriate to revisit the explication of the Ae. aegypti genome, as published in Science in 2007.1 This species is known to have radiated from the malaria vector Anopheles gambiae about 150 million years ago and the genome of that mosquito, along with the Drosophila melanogaster genome, was used for comparison purposes to understand the structure of the Aedes genome. The Aedes aegypti genome has a sequence size of 1.38 Gbp, 5-fold larger than Anopheles gambiae mosquito (278 Mbp); both mosquitos have three chromosomes but Ae. aegypti lacks the heteromorphic sex chromosomes found in D. melanogaster. The average gene length and size of intergenic regions was found to be 4-6 fold longer than in An. gambiae and D. melanogaster, which the authors believed was due to almost 50% of the Ae. aegypti genome comprising transposable elements. Despite these differences, chromosomal synteny was reported to have been maintained between these mosquito species, but orthologous gene order was found to be twice as high between the mosquito species than in comparison between either mosquito and the fruit fly.
Genetic analysis for the presence of open reading frames found 15,419 predicted proteins (and alternative splicing can increase this to 16,789 transcripts), with 80% of these confirmed as being transcriptionally active. There is an increased gene number for odorant binding proteins, cytochrome P450 and cuticle domains relative to An. gambiae. The relative distribution of genomic content was found as illustrated:
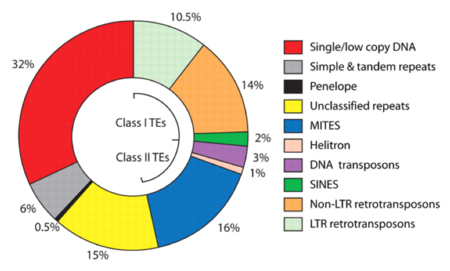 (where Penelope and Helitron are eukaryotic transposable elements and MITEs are miniature inverted repeat transposable elements that in Ae. aegypti have "terminal inverted repeat sequences and target-site duplications, features characteristic of transposition of DNA transposons" according to this report. These researchers identified over 1,000 types of transposable elements in the mosquitoe's genome, which included most of the families identified in eukaryotes, wherein non-protein coding transposable elements comprise 16% of the genome and are more numerous than protein-coding transposable elements (present at about 3% of the genome). Three percent of the Ae. aegypti genome comprise about 13,000 copies of long terminal repeat retrotransposons, and there are 50,000 copies of tRNA related SINE elements. The pattern of transposable element distribution in Ae. aegypti was found to be consistent with "an overall lack of apparent horizontal transfer events."
(where Penelope and Helitron are eukaryotic transposable elements and MITEs are miniature inverted repeat transposable elements that in Ae. aegypti have "terminal inverted repeat sequences and target-site duplications, features characteristic of transposition of DNA transposons" according to this report. These researchers identified over 1,000 types of transposable elements in the mosquitoe's genome, which included most of the families identified in eukaryotes, wherein non-protein coding transposable elements comprise 16% of the genome and are more numerous than protein-coding transposable elements (present at about 3% of the genome). Three percent of the Ae. aegypti genome comprise about 13,000 copies of long terminal repeat retrotransposons, and there are 50,000 copies of tRNA related SINE elements. The pattern of transposable element distribution in Ae. aegypti was found to be consistent with "an overall lack of apparent horizontal transfer events."
A consequence of the predominance of transposable elements in the Ae. aegypti genome created "an asymmetric distribution of intron lengths within genes" and frequent occurrence of transposable element-associated open reading frames "either in close proximity to genes or in introns" and an overall increase in gene length due to "infiltration" of transposable elements into introns. Compared to related species, "67% of the Ae. aegypti proteins having an ortholog in An. gambiae and 58% having an ortholog in D. melanogaster," with an "average amino acid identity of 74% between the mosquito proteins, in contrast with ~58% identity between mosquito and fruit fly proteins." Further:
Approximately 2,000 orthologs are shared only between the mosquitoes and may represent functions central to mosquito biology. Although most of these proteins are of unknown function, ~250 can be assigned a predicted function, of which 28% are involved in gustatory/olfactory systems, 12% are members of the cuticular gene family and 8% are members of the cytochrome P450 family.
These authors also report "an expansion of Zn-fingers, insect cuticle, cytochrome P450, odorant binding protein (OBP) A10/OS-D, insect allergen related and HMG-I and MHG-Y domains in Ae. aegypti" compared to other insects (specifically, An. gambiae, D. melanogaster and Apis mellifera (the honey bee)). Members of the superfamily of G-protein coupled receptors were also found, including "111 non-sensory class A, B and C GPCRs, 14 atypical class D GPCRs, and 10 opsin photoreceptors." Again in comparison with An. gambiae and D. melanogaster, these authors report orthologs for more than 85% of non-sensory GPCRs (which suggest to these researchers that there has been "significant conservation of GPCR-mediated neurological processes across [the] Diptera [family]." With regard to the ion transport-homologous genes found in the Aedes aegypti genome the paper states:
Similar to other multi-cellular eukaryotes, ~ 32% of all three insect transporters code for ion channels and probably function to maintain haemolymph homeostasis under different environmental conditions by modulating the concentrations of Na+, K+ and Cl− ions. Aedes aegypti encodes more paralogs of voltage-gated potassium ion channels, epithelial sodium channels and ligand-gated ion channels (LIC) such as the glutamate-gated ion channel than An. gambiae and D. melanogaster. These channels play important roles in the signal transduction pathway and cell communication in the central nervous system and at neuromuscular junctions. . . . A collection of 64 putative ATP-binding cassette transporters was identified, including subgroups that encode multi-drug efflux proteins. Aedes aegypti encodes more members of 4 different types of amino acid transporters than An. gambiae and D. melanogaster. Mosquito larvae cannot synthesize de novo all the basic, neutral or aromatic L-amino acids, and must rely on uptake of these essential amino acids. Aromatic amino acids phenylalanine and tryptophan are particularly important because they are precursors for the synthesis of neurotransmitters. The richer repertoire of membrane transport systems in Ae. aegypti is likely to intersect with the apparent increase in odorant reception and detoxification capacity.
Instead of bearing heteromorphic sex chromosomes, an Aedes aegypti gender is determined by an autosomal locus where the dominant allele determines maleness. The researchers detected gender differences in gene expression:
669 and 635 transcripts were enriched in females and males, respectively, and 6,713 transcripts were expressed at similar levels in both sexes.
An additional 373 and 534 transcripts generated exclusive hybridization signals (with signal intensity below cutoff threshold level in one channel) in females and males, respectively, and may therefore represent sex specific transcripts. Functional categorization of female and male enriched transcripts were remarkably similar, with male mosquitoes expressing a slightly larger number of immune and redox/stress related transcripts while females expressed a larger number of putative blood digestive enzyme transcripts. This particular pattern of immune gene expression is surprising considering a predicted lower need for immune defense in the males, due to the lack of pathogen exposure that results from blood feeding.
144 orthologous genes displaying the same sex specific transcription pattern in An. gambiae, while 74 orthologs showed an opposite profile, suggesting differences in certain sex specific functions between the two mosquito species.
Finally, the overall chromosomal organization showed significant synteny ("a distinct one-to-one correlation") between the Aedes aegypti genome and that of both An. gambiae and D. melanogaster (with an exception on the 2p and 3q chromosomal arms.
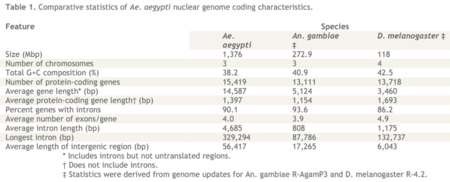
A summary of the genetic comparisons provided in the report was provided in tabular form as set forth below (click on table to enlarge).
1 This work was the result of an international effort involving scientists from The Institute for Genomic Research, Broad Institute of MIT and Harvard, Virginia Polytechnic Institute and State University, Bloomberg School of Public Health, Johns Hopkins University, University of Notre Dame, Harvard University, College of Agricultural Sciences, Colorado State University, Northwestern University, University of California, Riverside, University of California, Santa Cruz, Purdue University, Texas A&M, the Joint Technology Center, and the University of Massachusetts in the U.S.; European Bioinformatics Institute (EMBL-EBI), University of Geneva Medical School, Swiss Institute of Bioinformatics, Imperial College London, University of Oxford, Centro Nacional de Genotipado, Fundación Pública Galega de Medicina Xenómica, Hospital Clínico Universitario de Santiago, Institut Pasteur, Universidade da Coruña, Institut Jacques Monod, Universität Göttingen, and Complexo Hospitalario Universitario de Santiago in Europe; and Universidade de Sao Paulo, Instituto de Quimica, Universidade de Sao Paulo, Institute of Biomedical Sciences, and the Instituto Butantan in Brazil.

Leave a comment