By Kevin E. Noonan —

The duality of nucleic acids has long posed a challenge to patent law. A gene is, in one way of looking at it, "but a chemical compound, albeit a complex one," and this view of the gene's nature has informed the emphasis on "structure, structure, structure" taken by the Federal Circuit from Amgen Inc. v. Chugai Pharmaceutical Co. to Judge Rader's comment in the oral argument for In re Kubin. On the other hand, a gene also contains, and is defined by, the information it embodies, encoding an amino acid sequence corresponding to a structurally entirely different molecule, a protein. This aspect of a gene's nature, and the unpredictable variability of it, underlies both the increased stringency with which the written description requirement is applied to nucleic acid claims, as well as the Office's resistance to apply Judge Rader's "structure, structure, structure" rule in determining whether a novel nucleic acid sequence, identified using established methods, is non-obvious.
The latest example of this phenomenon is the Federal Circuit's decision last week to affirm rejection on 102/103 grounds of unpatentability in In re Greave. The claims at issue include this generic claim:
(where IGFBP-2 and -5 stand for two different species of insulin growth factor binding protein). Also among the rejected claims were claims directed to particular species as set forth in applicants' specification. The prior art asserted in support of the rejection was a PCT Publication to Wraight (WO 00/78431), which disclosed a list of all 15-mer oligonucleotides that could be made from the (publicly-available) full-length IGFBP-2 sequence, as well as disclosure in the Wraight reference that antisense oligonucleotides could comprise any oligonucleotide that specifically could bind to (i.e., hybridize with) any of the disclosed 15-mer sense oligonucleotides, and the existence in the Wraight disclosure of several 15-mer sense oligonucleotides with sequence complimentarity overlap to the specific oligonucleotides disclosed and claimed by Gleave.
The Federal Circuit affirmed the rejection, in a decision by Judge Prost joined by Chief Judge Michel and Judge Moore. The Court based this decision on its view that anticipation required merely that the oligonucleotide sequence was in the prior art, not that its usefulness was previously disclosed. Gleave argued that the art disclosed a mere list comprising "ink," not disclosure of any functional antisense oligonucleotide (indeed, the cited art contained no disclosure of an antisense oligonucleotide at all), but such disclosure was unnecessary, according to the Court. All that is necessary for a prior art reference to anticipate, said the Court, is that it enable the skilled artisan to make the claimed invention, not use it, citing Impax Lab., Inc. v. Aventis Pharm. Inc., 545 F.3d 1312, 1314 (Fed. Cir. 2008); and In re LeGrice, 301 F.2d 929, 940–44 (C.C.P.A. 1962). The prior art disclosure need not show actual reduction to practice (Schering Corp. v. Geneva Pharm., Inc., 339 F.3d 1373, 1380–81 (Fed. Cir. 2003); In re Donohue, 766 F.2d 531, 533 (Fed. Cir. 1985)), and the description provided in the putatively anticipating reference "might not otherwise entitle its author to a patent," citing Vas-Cath Inc. v. Mahurkar, 935 F.2d 1555, 1562 (Fed. Cir. 1991).
Cases that could be interpreted to the contrary, such as Bristol-Myers Squibb Co. v. Ben Venue Lab., Inc., 246 F.3d 1368, 1374 (Fed. Cir. 2001) ("[t]o anticipate, the reference must also enable one of skill in the art to make and use the claimed invention") are "[t]aken out of context," according to the Court, insofar as they are interpreted to impose an operability or utility disclosure requirement for an anticipating reference. A "thorough" reading of the Court's caselaw (including Novo Nordisk Pharm., Inc. v. Bio-Technology General Corp., 424 F.3d 1347, 1355 (Fed. Cir. 2005); Rasmusson v. SmithKline Beecham Corp., 413 F.3d 1318, 1326 (Fed. Cir. 2005); and In re Hafner, 410 F.2d 1403, 1405 (C.C.P.A. 1969)) imposes no such requirement, the Court concluded. And the Court also noted that Gleave's claims themselves contained no limitation restricting their scope to functional antisense oligonucleotides.
The crux of the distinction the Court made, regarding the use of a list of sense oligonucleotides in the cited art and the traditional proscription against using a mere recitation of all possible compounds as anticipating claims to particular species, is found in the Court's consideration of the application of In re Wiggins, 488 F.2d 538, 543 (C.C.P.A. 1973) to the facts before it. In Wiggins, the Court of Customs and Patent Appeals stated:
If we were to hold otherwise, lists of thousands of theoretically possible compounds could be generated and published which, assuming it would be within the level of skill in the art to make them, would bar a patent to the actual discoverer of a named compound no matter how beneficial to mankind it might be.
488 F.2d at 543 (emphases added).
The Court noted that the CCPA in Wiggins stated that "'[t]he mere naming of a compound in a reference, without more, cannot constitute a description of the compound.'" 488 F.2d at 543. But the distinction with this case, according to the Court's opinion, is that Wiggins involved the "mere naming of a theoretical compound, without more," and that was not anticipating. Indeed, the Court emphasized that "'[w]ithout more' is the key phrase," because what is "more" is the capacity for the person of ordinary skill in the art to be able to make the claimed compound. Here, these were not "potential or theoretical" compounds, but a class of compounds that fell within so limited a genus that the skilled artisan could "'at once envision each member of this limited class,'" citing Eli Lilly & Co. v. Zenith Goldline Pharm., Inc., 471 F.3d 1369, 1376 (Fed. Cir. 2006). "In that limited circumstance, a reference describing the genus anticipates every species within the genus. See Perricone [v. Medicis Pharm. Corp., 432 F.3d 1368, 1377 (Fed. Cir. 2005)]."
What was not anticipated by the cited art are methods of using these oligonucleotides, and the Court suggested that Gleave might be entitled to such claims. As for the appealed composition of matter claims, the Court found that these sequences "are simply not new," and affirmed. And here is where the duality of nucleic acids becomes important. It was undisputed that the Wraight reference did not disclose a single antisense oligonucleotide — it merely disclosed all 15-mer sense oligonucleotides, coupled with generic disclosure that antisense oligonucleotides would specifically hybridize to the disclosed sense oligonucleotides, and the knowledge in the art that such oligonucleotides could be made without undue experimentation. Regardless of operability (although Gleave's arguments might have had additional force had they contained a functional limitation regarding operability as antisense oligonucleotides), the Court's decision suggests that nucleic acids can be anticipated by the mere recitation of any sequence, or its complement, in the prior art regardless of any disclosure as to operability. This raises the question of whether an unidentified open reading frame, published in a publicly-available database, might not be enough to anticipate claims directed to an isolated nucleic acid encoding a novel protein. Like the antisense oligonucleotides at issue in Gleave, the existence of the sequence in the prior art could be combined with the knowledge of the skilled worker that proteins are encoded by open reading frames to anticipate the sequence. Known operability is not required, according to this panel opinion, raising the question of whether the availability of the published but unrecognized sequence would be enough. There are certainly distinctions: here, the oligonucleotides comprised a known protein, for example. And there would be an In re Hall question about whether an unannotated open reading frame would be sufficiently accessible to qualify as prior art. But the Federal Circuit's decision in Gleave amounts to yet another instance of differential applicatio
n of the law to biotechnology claims, whose effects remain to be fully appreciated.
Panel: Chief Judge Michel and Circuit Judges Prost and Moore
Opinion by Circuit Judge Prost

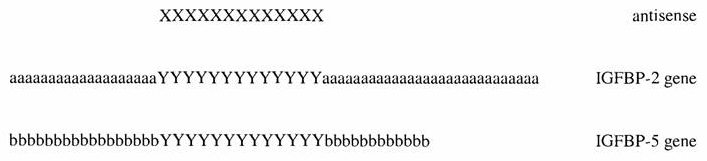
Leave a comment